5. Project Models to a Single Scenario
Source:vignettes/model_predictions.Rmd
model_predictions.RmdSummary
- Description
- Getting ready
- Fitted models
- Model predictions
- Options for predictions
- Binarize predictions
- Saving Predictions
- Detecting changes in predictions
Description
Once selected models have been fit and explored, projections to
single or multiple scenarios can be performed. The
predict_selected() function is designed for projections to
single scenarios (i.e., a single set of new data). This vignette
contains examples of how to use many of the options available for model
predictions.
Getting ready
At this point it is assumed that kuenm2 is installed (if
not, see the Main
guide). Load kuenm2 and any other required packages,
and define a working directory (if needed).
Note: functions from other packages (i.e., not from base R or
kuenm2) used in this guide will be displayed as
package::function().
# Load packages
library(kuenm2)
library(terra)
# Current directory
getwd()
# Define new directory
#setwd("YOUR/DIRECTORY") # uncomment and modify if setting a new directory
# Saving original plotting parameters
original_par <- par(no.readonly = TRUE)Fitted models
To predict using the selected models, a fitted_models
object is required. For detailed information on model fitting, check the
vignette Fit and Explore Selected
Models. The fitted_models object generated in that
vignette is included as an example dataset within the package. Let’s
load it.
# Import fitted_model_maxnet
data("fitted_model_maxnet", package = "kuenm2")
# Print fitted models
fitted_model_maxnet
#> fitted_models object summary
#> ============================
#> Species: Myrcia hatschbachii
#> Algortihm: maxnet
#> Number of fitted models: 2
#> Models fitted with 4 replicatesTo compare the results, let’s import a fitted_models
object generated using the GLM algorithm:
# Import fitted_model_glm
data("fitted_model_glm", package = "kuenm2")
# Print fitted models
fitted_model_glm
#> fitted_models object summary
#> ============================
#> Species: Myrcia hatschbachii
#> Algortihm: glm
#> Number of fitted models: 1
#> Only full models fitted, no replicatesModel predictions
To predict selected models for a single scenario, you need a
fitted_models object and the corresponding variables. These
variables can be provided as either a SpatRaster or a
data.frame. The names of the variables (or columns in the
data.frame) must precisely match those used for model
calibration or those used when running PCA (if
do_pca = TRUE was set in the prepare_data()
function; see Prepare Data for Model
Calibration for more details).
Predict to SpatRaster
Let’s use the same raster variables that were used to prepare the data and calibrate the models. These are included as example data within the package:
# Import raster layers
var <- rast(system.file("extdata", "Current_variables.tif", package = "kuenm2"))
# Plot raster layers
terra::plot(var)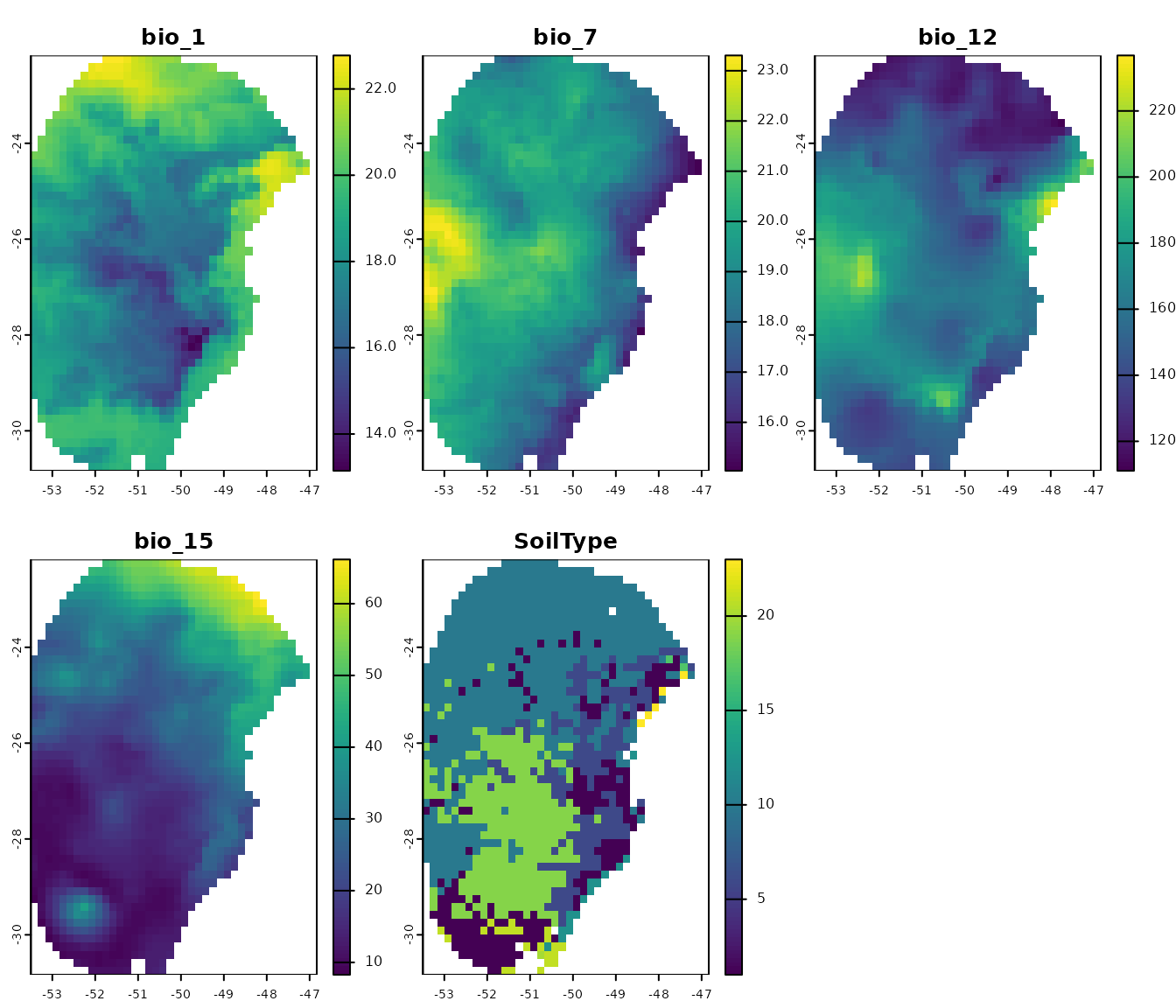
Let’s check which variables were used to calibrate our models. They
are available in the calibration_data element of the
object:
# Variables used to calibrate maxnet models
colnames(fitted_model_maxnet$calibration_data)
#> [1] "pr_bg" "bio_1" "bio_7" "bio_12" "bio_15" "SoilType"
# Variables used to calibrate glms
colnames(fitted_model_glm$calibration_data)
#> [1] "pr_bg" "bio_1" "bio_7" "bio_12" "bio_15" "SoilType"The first column, “pr_bg”, indicates the presence (1) and background
(0) records, while the other columns represent the environmental
variables. In this case, the variables are bio_1,
bio_7, bio_12, bio_15, and
SoilType. All these variables are present in the
SpatRaster (var) imported, so, we can predict
our models to this raster. Let’s begin by predicting the maxnet
model:
p_maxnet <- predict_selected(models = fitted_model_maxnet, new_variables = var,
progress_bar = FALSE)By default, the function computes consensus metrics (mean, median,
range, and standard deviation) for each model across its replicates (if
they were produced), as well as a general consensus across all models
(if multiple were selected). In this case, the output is a
list containing SpatRasters for predictions,
the consensus for each model, and the general consensus:
# See objects in the output of predict_selected
names(p_maxnet)
#> [1] "Model_192" "Model_219" "General_consensus"Let’s plot the general consensus:
terra::plot(p_maxnet$General_consensus)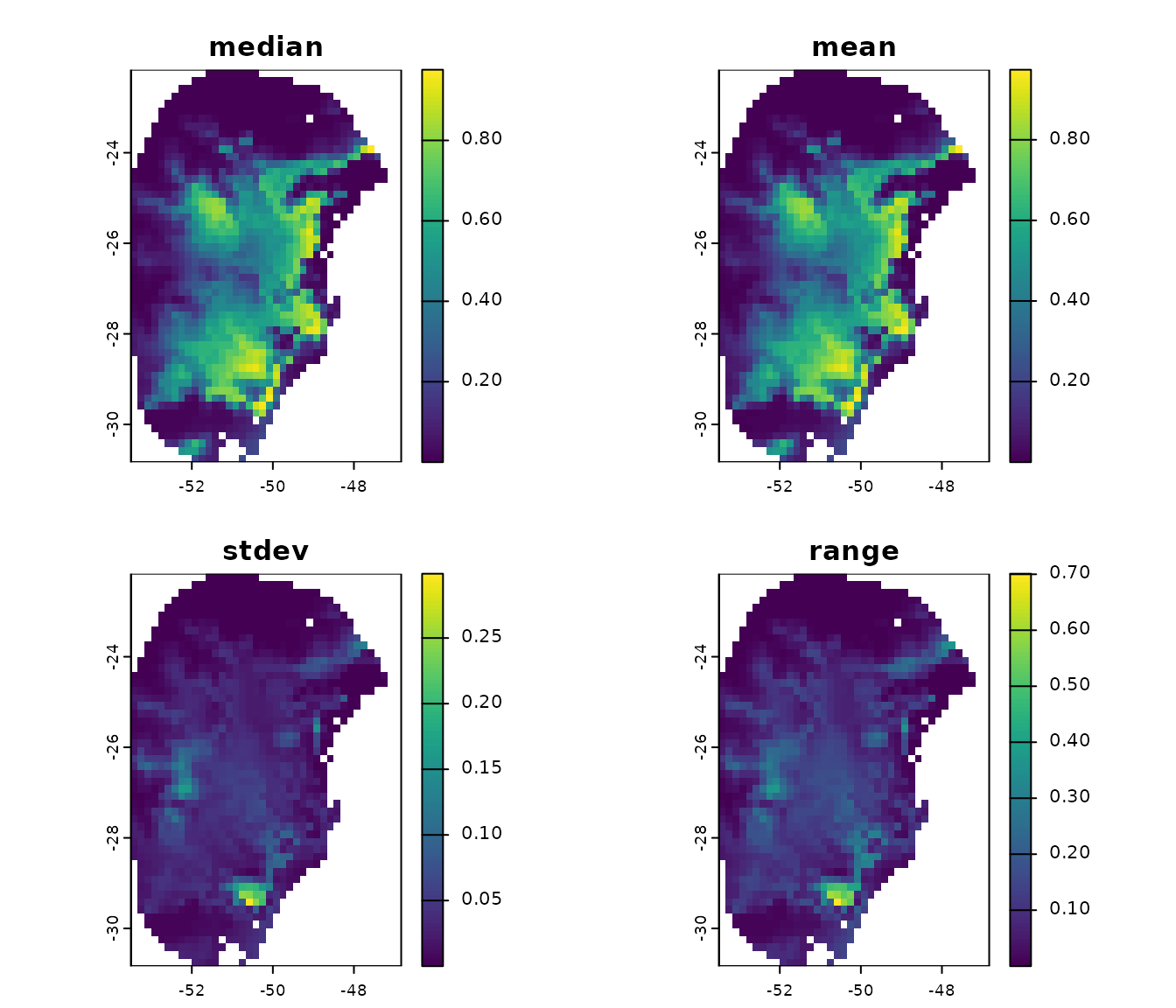
We can also plot the results for each replicate and the consensus for each model:
# Predictions for each replicate from model 192
terra::plot(p_maxnet$Model_192$Replicates)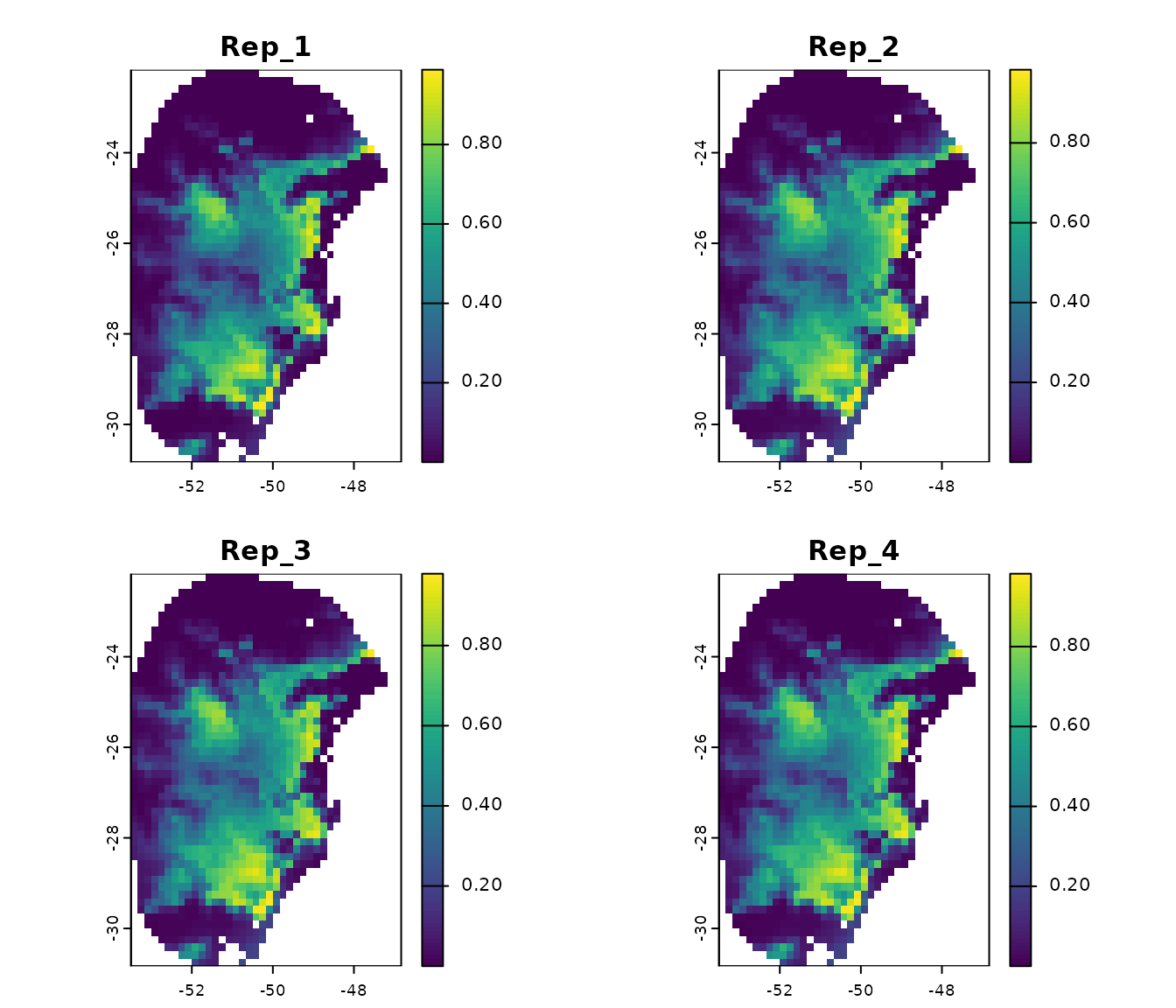
# Consensus across each replicate from model 192
terra::plot(p_maxnet$Model_192$Model_consensus)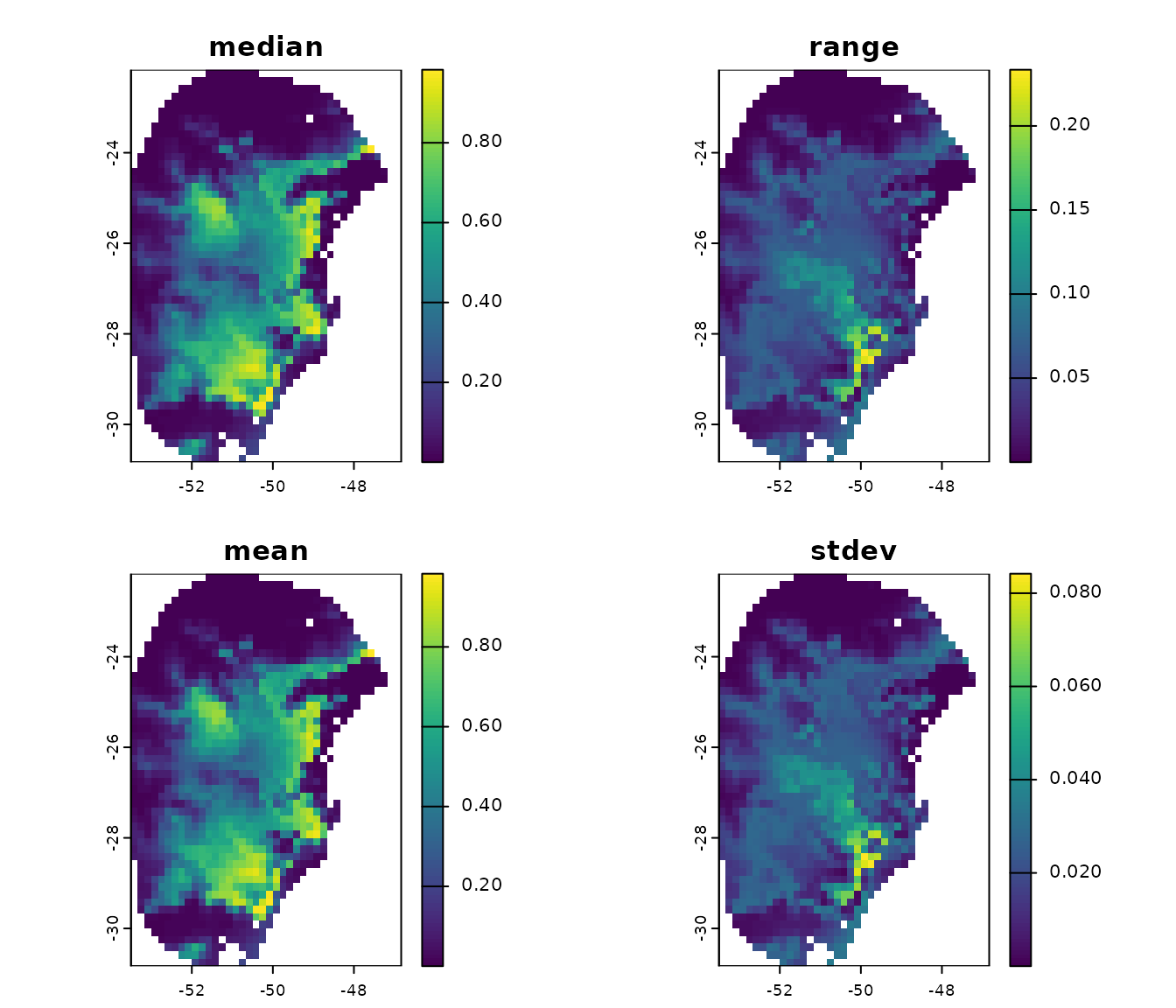
For comparison, let’s predict the GLM:
# Predict glm
p_glm <- predict_selected(models = fitted_model_glm,
new_variables = var,
progress_bar = FALSE)
# See selected models that were predicted
names(p_glm)
#> [1] "Model_85" "General_consensus"
# Compare general consensus (mean) between maxnet and glm
par(mfrow = c(1, 2)) # Set grid to plot
terra::plot(p_maxnet$General_consensus$mean, main = "Maxnet")
terra::plot(p_glm$General_consensus, main = "GLM")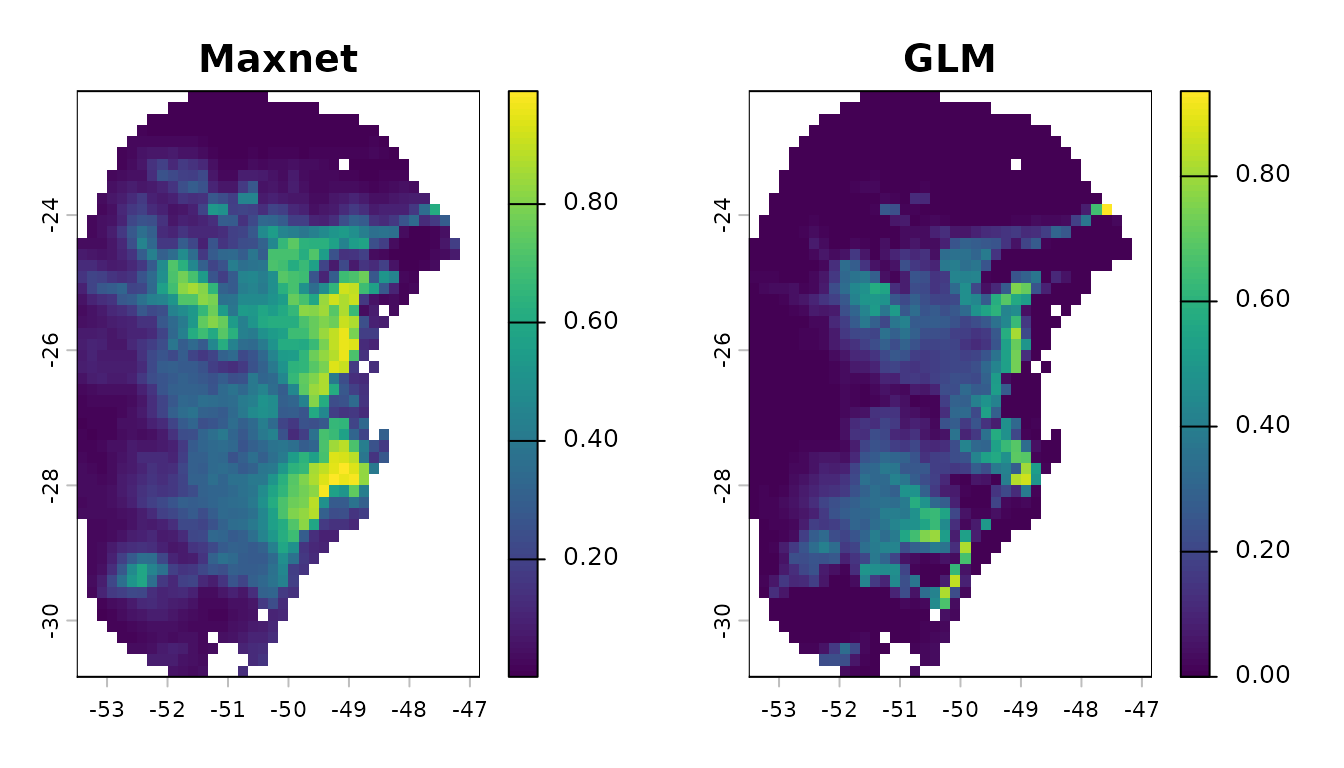
Predict to data.frame
Instead of a SpatRaster, we can also predict the models
to a data.frame with the variable values. As an example,
let’s convert the raster variables var to a
data.frame:
var_df <- as.data.frame(var)
head(var_df)
#> bio_1 bio_7 bio_12 bio_15 SoilType
#> 11 22.77717 18.12400 1180 48.03594 NA
#> 12 22.76711 17.74400 1191 49.31194 10
#> 13 22.68580 17.46575 1206 51.51922 10
#> 14 22.50121 17.84525 1228 53.90265 10
#> 15 22.07609 18.14125 1254 54.10397 10
#> 16 21.88485 18.80800 1276 54.07279 10Note that each column stores the values for each variable. Let’s
predict our Maxnet models to this data.frame:
p_df <- predict_selected(models = fitted_model_maxnet,
new_variables = var_df, # Now, a data.frame
progress_bar = FALSE) Now, instead of SpatRaster objects, the function returns
data.frame objects with the predictions:
# Results by replicate of the model 192
head(p_df$Model_192$Replicates)
#> Replicate_1 Replicate_2 Replicate_3 Replicate_4
#> 1 0.006521501 0.0006209852 0.0005883615 9.831561e-05
#> 2 0.006446437 0.0005356316 0.0005713501 9.009486e-05
#> 3 0.006233583 0.0003396879 0.0004975279 6.967025e-05
#> 4 0.005797668 0.0001458500 0.0003605775 4.303576e-05
#> 5 0.008513515 0.0002034105 0.0006532983 8.529550e-05
#> 6 0.009240381 0.0001753492 0.0006784553 9.035171e-05
# Consensus across replicates of the model 192
head(p_df$Model_192$Model_consensus)
#> median range mean stdev
#> 1 0.0006046734 0.006423186 0.001957291 0.003052184
#> 2 0.0005534909 0.006356342 0.001910878 0.003031621
#> 3 0.0004186079 0.006163912 0.001785117 0.002970901
#> 4 0.0002532137 0.005754632 0.001586783 0.002810372
#> 5 0.0004283544 0.008428219 0.002363880 0.004107054
#> 6 0.0004269022 0.009150029 0.002546134 0.004470371
# General consensus across all models
head(p_df$General_consensus)
#> median range mean stdev
#> 1 0.0006049792 0.006423186 0.001882943 0.002691096
#> 2 0.0005534909 0.006356342 0.001847258 0.002690840
#> 3 0.0004186079 0.006163912 0.001737935 0.002664381
#> 4 0.0002532730 0.005757458 0.001562733 0.002558575
#> 5 0.0004283544 0.008435158 0.002337815 0.003756587
#> 6 0.0004274250 0.009165160 0.002539356 0.004130584Options for predictions
Output type
Maxnet models produce four different types of output for their predictions: raw, cumulative, logistic, and cloglog. These are described in Merow et al. 2013 and Phillips et al. 2017.
All four output types are monotonically related; thus, rank-based metrics for model fit (e.g., omission rate and partial ROC) will be identical. However, the output types have different scaling, which leads to distinct interpretations and visually different prediction maps.
- Raw (or exponential) output is interpreted as a Relative Occurrence Rate (ROR). The ROR sums to 1 when predicted to the data used to train the model.
- Cumulative output assigns to a location the sum of all raw values less than or equal to the raw value for that location, and then rescales this to range between 0 and 100. Cumulative output can be interpreted in terms of an omission rate because thresholding at a value of c to predict a suitable/unsuitable cell will omit approximately c% of presences.
- Cloglog output (Default) transforms the raw values into a scale of relative suitability ranging between 0 and 1, using a logistic transformation based on a user-specified parameter ‘’, which represents the probability of presence at ‘average’ presence locations. In this context, the tau value defaults to .
- Logistic output is similar to Cloglog, but it assumes that .
Let’s examine the differences between these four output types for Maxnet models:
p_cloglog <- predict_selected(models = fitted_model_maxnet, new_variables = var,
type = "cloglog", progress_bar = FALSE)
p_logistic <- predict_selected(models = fitted_model_maxnet, new_variables = var,
type = "logistic", progress_bar = FALSE)
p_cumulative <- predict_selected(models = fitted_model_maxnet, new_variables = var,
type = "cumulative", progress_bar = FALSE)
p_raw <- predict_selected(models = fitted_model_maxnet, new_variables = var,
type = "raw", progress_bar = FALSE)
# Plot the differences
par(mfrow = c(2, 2))
terra::plot(p_cloglog$General_consensus$mean, main = "Cloglog (Default)",
zlim = c(0, 1))
terra::plot(p_logistic$General_consensus$mean, main = "Logistic",
zlim = c(0, 1))
terra::plot(p_cumulative$General_consensus$mean, main = "Cumulative",
zlim = c(0, 1))
terra::plot(p_raw$General_consensus$mean, main = "Raw", zlim = c(0, 1))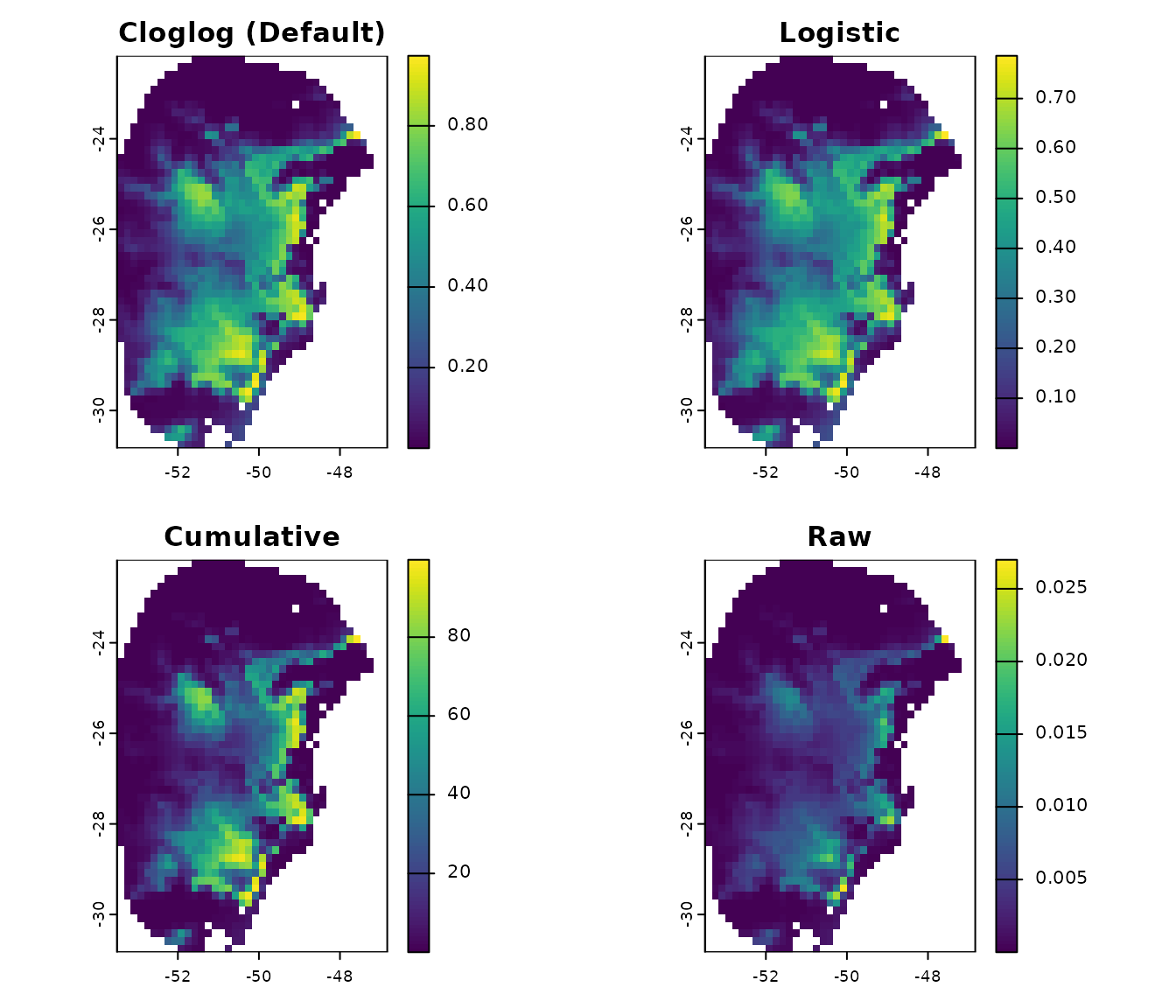
Clamping variables
By default, predictions are performed with free extrapolation
(extrapolation_type = "E"). This can be problematic when
the peak of suitability occurs at the extremes of a variable’s range.
For example, let’s examine the response curve of the Maxnet model for
bio_7 (Temperature Annual Range):
response_curve(models = fitted_model_maxnet, variable = "bio_7",
extrapolation_factor = 1)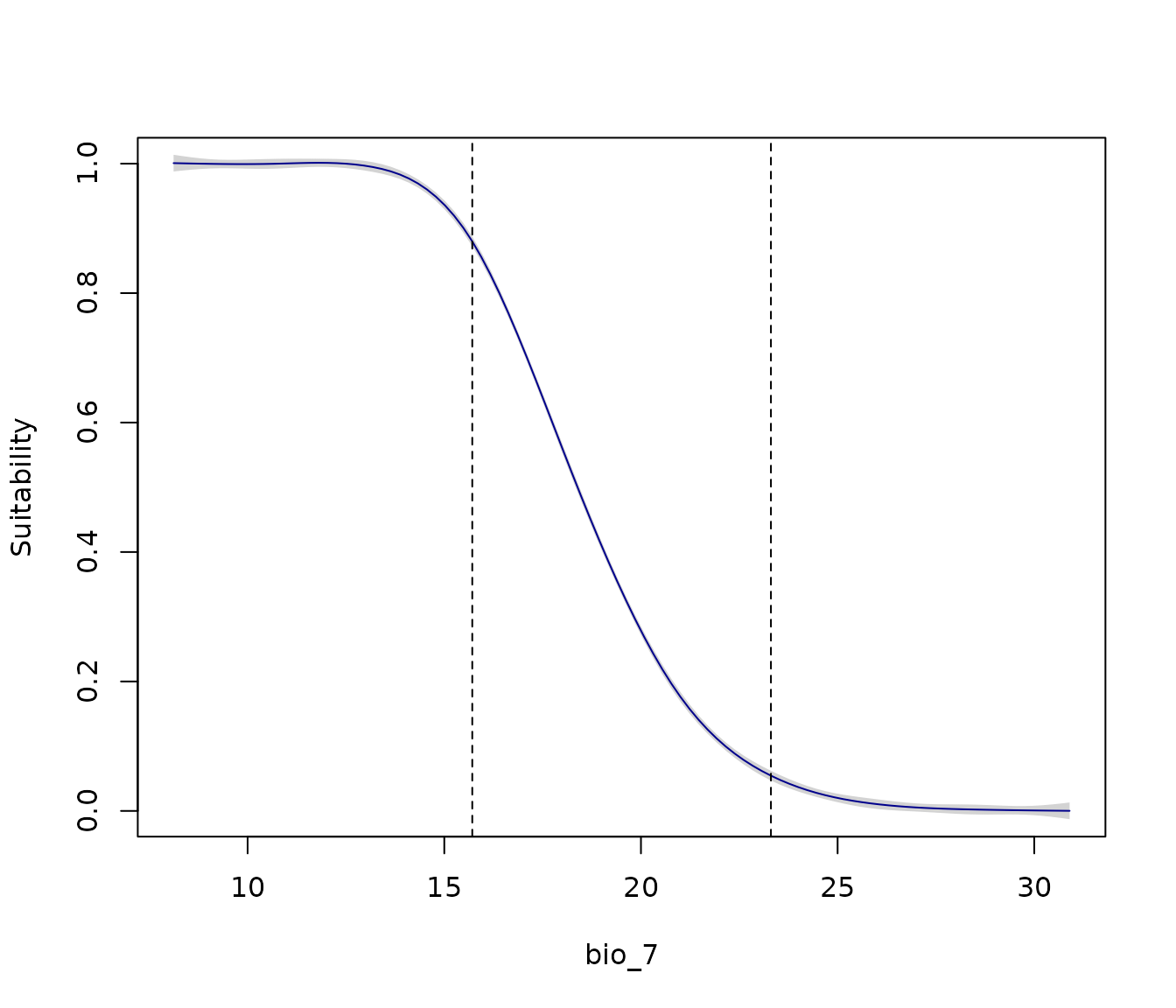
Note that higher suitability occurs at low values of the temperature
range. However, the lower limit of the calibration data used to fit the
models (dashed line) is at 15.7ºC. The premise that suitability will
increase and stabilize at lower values of bio_7 is an
extrapolation of the model (the area to the left of the dashed line).
It’s possible that suitability decreases at extremely low values, but
training data is insufficient for the model to predict this.
One way to address this is by clamping the variables. This means that
all prediction values outside the training range (both below the lower
value and above the upper value) are set to the prediction values found
at the limits of the range. For example, in the calibration data for the
Maxnet models, the lower and upper limits for bio_7 are
15.7ºC and 23.3ºC, respectively:
range(fitted_model_maxnet$calibration_data$bio_7)
#> [1] 15.71120 23.30475To observe the effect of clamping this variable, let’s create a
hypothetical scenario where bio_7 has very low values:
# From bio_7, reduce values
new_bio7 <- var$bio_7 - 3
# Create new scenario
new_var <- var
# Replace bio_7 with new_bio7 in this scenario
new_var$bio_7 <- new_bio7
# Plot the differences
par(mfrow = c(1, 2))
terra::plot(var$bio_7, main = "Original bio_7", range = c(5, 25))
terra::plot(new_var$bio_7, main = "New bio_7", range = c(5, 25))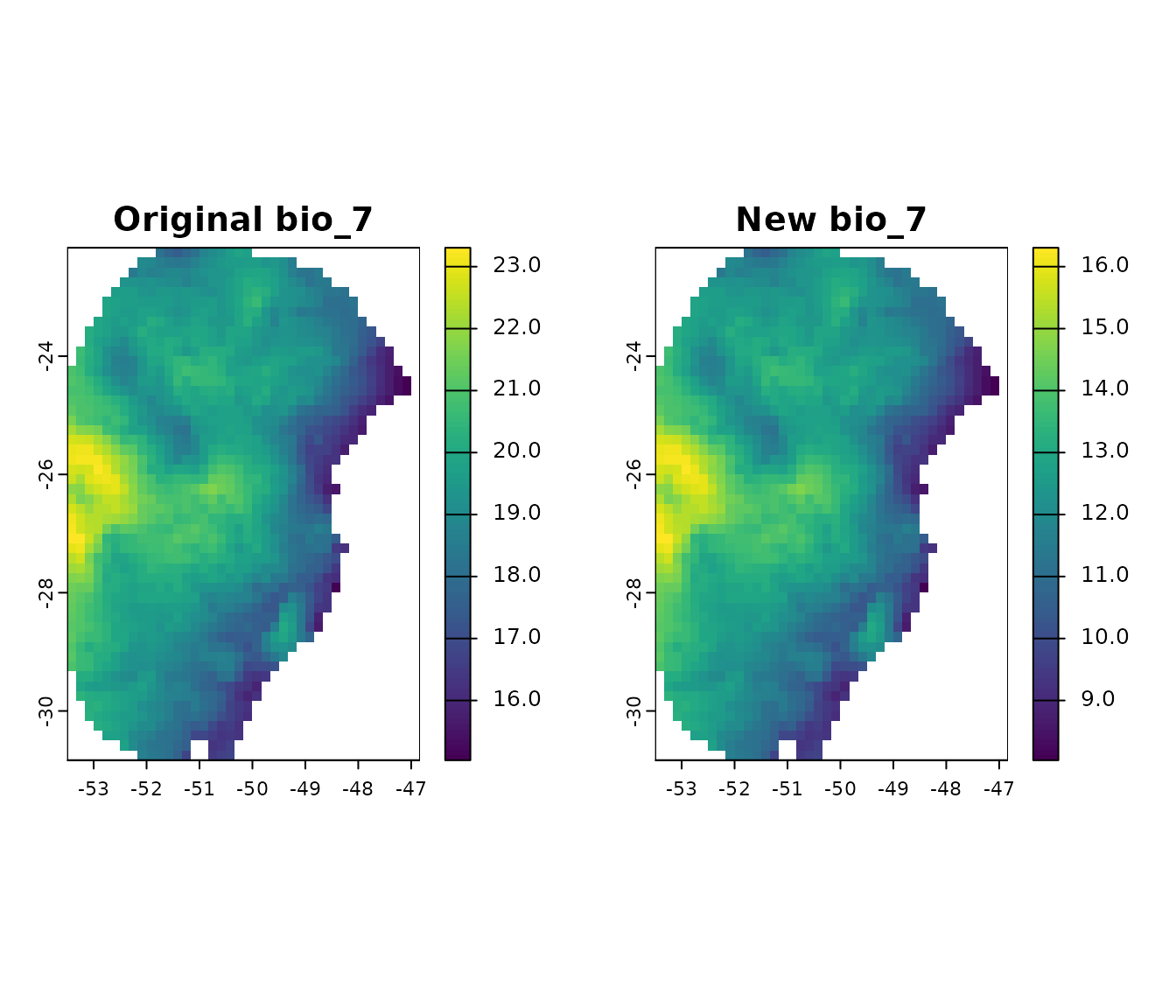
Let’s predict the Maxnet models for this new scenario with both free
extrapolation (extrapolation_type = "E") and with clamped
variables (extrapolation_type = "EC"):
# Predict to hypothetical scenario with free extrapolation
p_free_extrapolation <- predict_selected(models = fitted_model_maxnet,
new_variables = new_var, # New scenario
consensus = "mean",
extrapolation_type = "E", # Free extrapolation (Default)
progress_bar = FALSE)
# Predict to hypothetical scenario with clamping
p_clamping <- predict_selected(models = fitted_model_maxnet,
new_variables = new_var, # New scenario
consensus = "mean",
extrapolation_type = "EC", # Extrapolation with clamping
progress_bar = FALSE)
# Get and see differences
p_difference <- p_free_extrapolation$General_consensus$mean - p_clamping$General_consensus$mean
# Plot the differences
par(mfrow = c(2, 2))
terra::plot(p_free_extrapolation$General_consensus$mean,
main = "Free extrapolation", zlim = c(0, 1))
terra::plot(p_clamping$General_consensus$mean, main = "Clamping",
zlim = c(0, 1))
terra::plot(p_difference, main = "Difference")
terra::plot(new_bio7, main = "Hypothetical bio_7", type = "interval")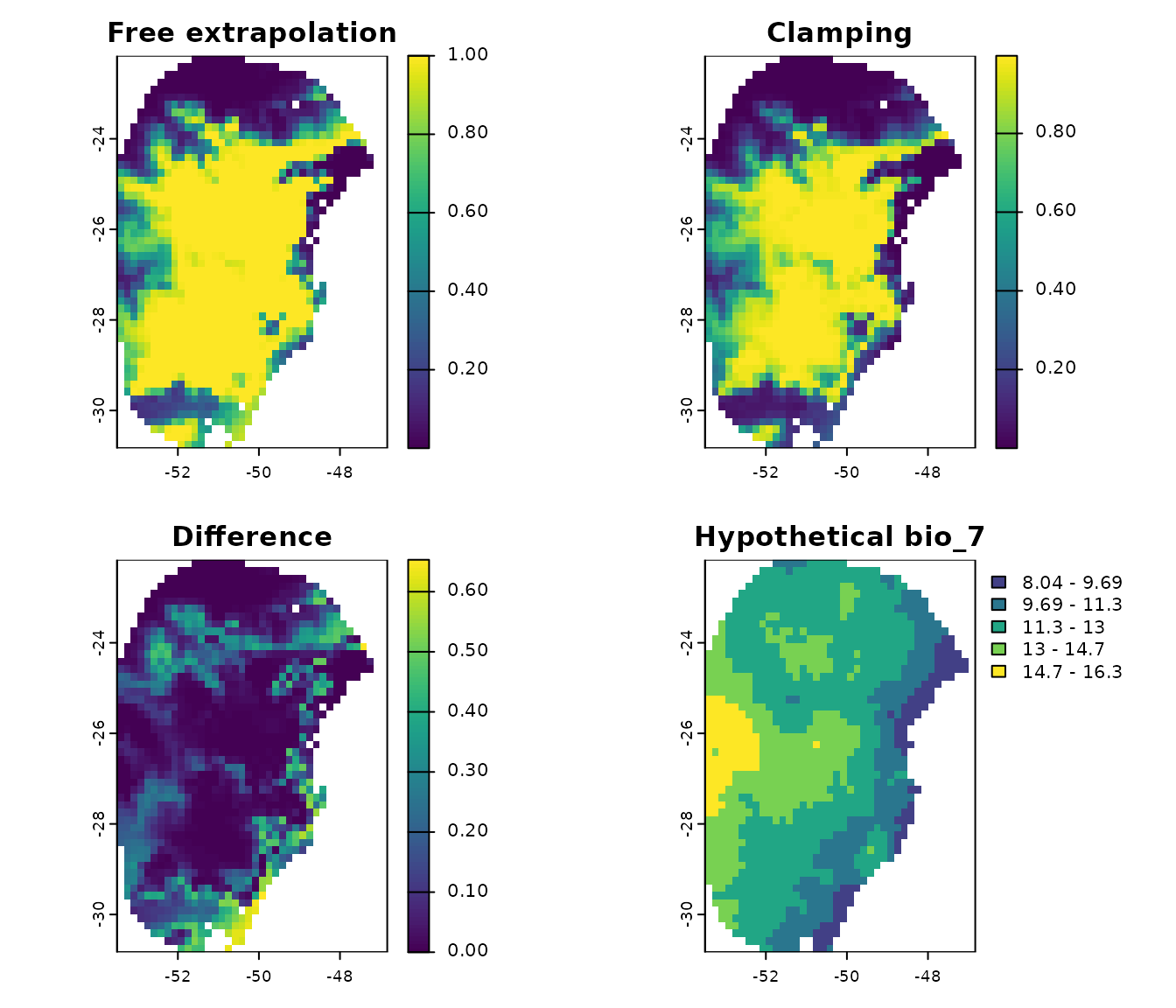
Note that when we clamp the variables, regions with extremely low
values of (the hypothetical) bio_7 exhibit lower predicted
suitability values compared to when free extrapolation is allowed.
By default, when extrapolation_type = "EC" is set, all
variables are clamped. You can specify which variables to clamp using
the var_to_restrict argument.
No extrapolation
A more rigorous approach is to predict with no extrapolation. Here regions outside the limits of the training data are assigned a suitability value of 0. Let’s proceed to observe the differences:
# Predict to hypothetical scenario with no extrapolation
p_no_extrapolation <- predict_selected(models = fitted_model_maxnet,
new_variables = new_var, # New scenario
consensus = "mean",
extrapolation_type = "NE", # No extrapolation
progress_bar = FALSE)
# Plot the differences
par(mfrow = c(2, 2))
terra::plot(p_free_extrapolation$General_consensus$mean, main = "Free extrapolation",
zlim = c(0, 1))
terra::plot(p_clamping$General_consensus$mean, main = "Clamping",
zlim = c(0, 1))
terra::plot(p_no_extrapolation$General_consensus$mean, main = "No extrapolation",
zlim = c(0, 1))
terra::plot(new_bio7, main = "Hypothetical bio_7", type = "interval")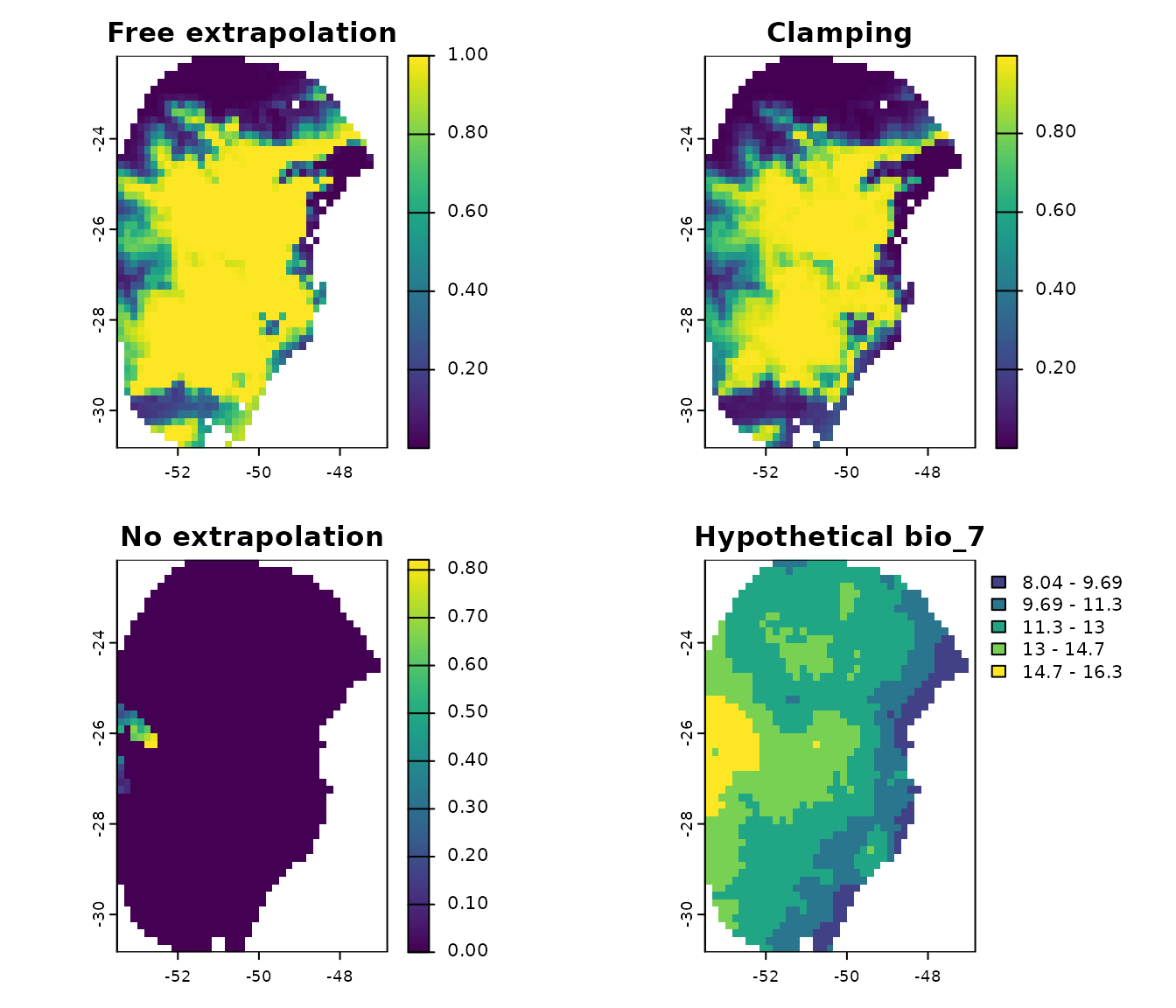
In this example, a large portion of the predicted area shows zero
suitability. This is because, in this hypothetical scenario, much of the
region has bio_7 values lower than those in the training
data, which has a minimum of 15ºC. Suitability values greater than zero
are only in areas where bio_7 falls within the training
range.
By default, when extrapolation_type = "NE" is set, all
variables are considered for this process. You can specify a subset of
variables to be considered for extrapolation using the
var_to_restrict argument.
Binarize predictions
The fitted_models object stores the thresholds that can
be used to classify model predictions into suitable and unsuitable
areas. These thresholds correspond to the omission error rate used
during model selection (e.g., 5% or 10%).
You can access the omission error rate used to calculate the thresholds directly from the object:
# Get omission error used to select models and calculate the thesholds
## For maxnet model
fitted_model_maxnet$omission_rate
#> [1] 10
## For glm
fitted_model_glm$omission_rate
#> [1] 10In both models, a 10% omission error rate was used to calculate the thresholds. This means that when predictions are binarized, approximately 10% of the presence records used to train models will fall into areas classified as unsuitable.
The thresholds are summarized in two ways: the mean and median across replicates for each model, and the consensus mean and median across all selected models (when more than one model is selected). Let’s check the thresholds for the general consensus:
# For maxnet
fitted_model_maxnet$thresholds$consensus
#> $mean
#> [1] 0.3095083
#>
#> $median
#> [1] 0.259534
# For glm
fitted_model_glm$thresholds$consensus
#> $mean
#> [1] 0.1204713
#>
#> $median
#> [1] 0.1204713Let’s use these threshold values to binarize models predictions:
# Get the threshold values for models (general consensus)
thr_mean_maxnet <- fitted_model_maxnet$thresholds$consensus$mean # Maxnet
thr_mean_glm <- fitted_model_glm$thresholds$consensus$mean # glm
# Binarize models
mean_maxnet_bin <- (p_maxnet$General_consensus$mean >= thr_mean_maxnet) * 1
mean_glm_bin <- (p_glm$General_consensus >= thr_mean_glm) * 1
# Compare results
par(mfrow = c(1, 2)) # Set grid to plot
terra::plot(mean_maxnet_bin, main = "Maxnet")
terra::plot(mean_glm_bin, main = "GLM")
# Reset plotting parameters
par(original_par) Saving predictions
We can save the predictions to the disk by setting
write_files = TRUE. When this option is enabled, you must
provide a directory path in the out_dir argument.
If new_variables is a SpatRaster, the
function will save files as GeoTIFF (.tif) files. If
new_variables is a data.frame, the function
will save the output files as Comma Separated Value (.csv) files.
p_save <- predict_selected(models = fitted_model_maxnet,
new_variables = var,
write_files = TRUE, # To save to the disk
write_replicates = TRUE, # To save predictions for each replicate
out_dir = tempdir(), # Directory to save the results (temporary directory)
progress_bar = FALSE)Alternatively, we can use writeRaster() to save specific
output predictions manually. For example, to save only the mean layer
from the general consensus results:
terra::writeRaster(p_maxnet$General_consensus$mean,
filename = file.path(tempdir(), "Mean_consensus.tif"))Detecting changes in predictions
To compare predictions between two single scenarios representing
different time periods (e.g., present vs. future or present vs. past),
the function prediction_changes() can be used. This
function helps to identify loss (contraction),
gain (expansion), and stability (no
change) of suitable areas.
As an example, we will project the fitted model to a single GCM representing future climatic conditions:
# Read layers representing future conditions
future_var <- terra::rast(system.file("extdata",
"wc2.1_10m_bioc_ACCESS-CM2_ssp585_2081-2100.tif",
package = "kuenm2"))
# Plot future layers
terra::plot(future_var)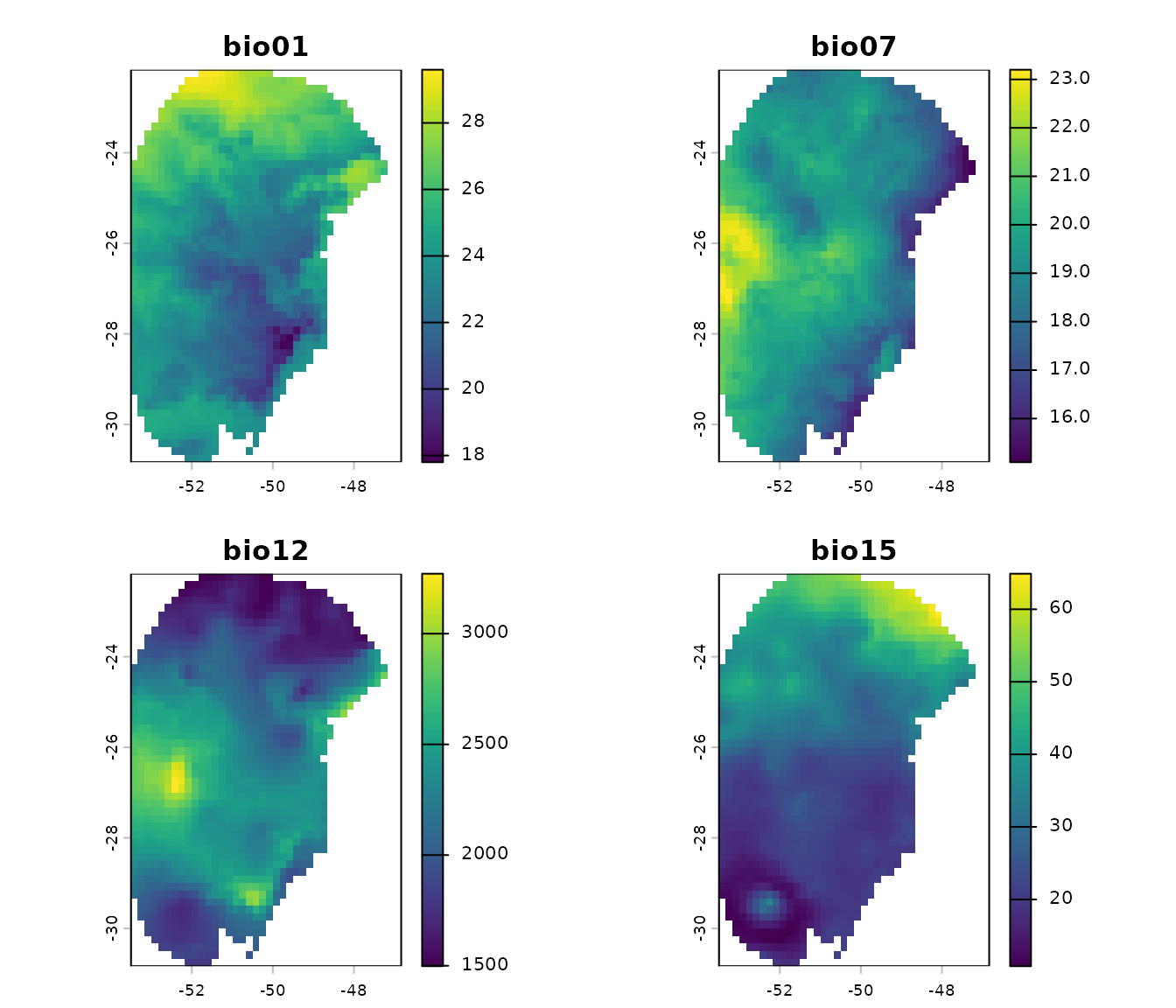
Next, we need to rename the variables so that they match the variable names used when fitting the models. After that, we will also append the static soil variable to the set of future variables.
# renaming layers to match names of variables used to fit the model
names(future_var) <- sub("bio0", "bio", names(future_var))
names(future_var) <- sub("bio", "bio_", names(future_var))
names(var)
#> [1] "bio_1" "bio_7" "bio_12" "bio_15" "SoilType"
names(future_var)
#> [1] "bio_1" "bio_7" "bio_12" "bio_15"
# Adding soil layer to future variable set
future_var <- c(future_var, var$SoilType)Now we can generate predictions under future environmental conditions:
# Predict
p_future <- predict_selected(models = fitted_model_maxnet,
new_variables = future_var,
progress_bar = FALSE)
# Plot consensus (mean)
terra::plot(c(p_maxnet$General_consensus$mean,
p_future$General_consensus$mean),
main = c("Present", "Future (SSP 585, 2081-2100)"))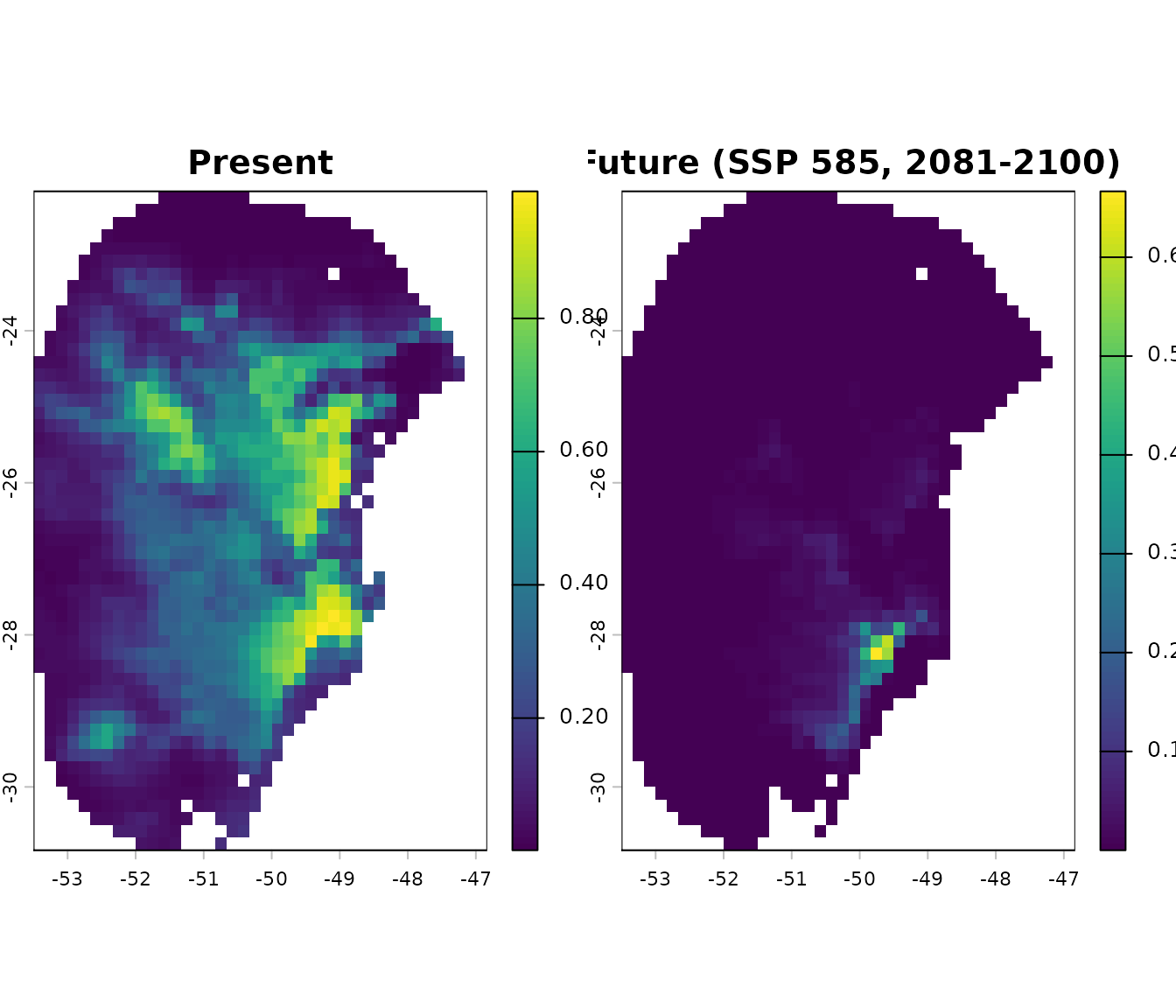
To identify how suitable areas change between scenarios, we can use
prediction_changes(). This function computes binary layers
from the predictions using the threshold stored in the fitted models,
compares current and future predictions, and then classifies each cell
as gain, loss, or stable.
# Compute changes between scenarios
p_changes <- prediction_changes(current_predictions = p_maxnet$General_consensus$mean,
new_predictions = p_future$General_consensus$mean,
fitted_models = fitted_model_maxnet,
predicted_to = "future")
# Plot result
terra::plot(p_changes)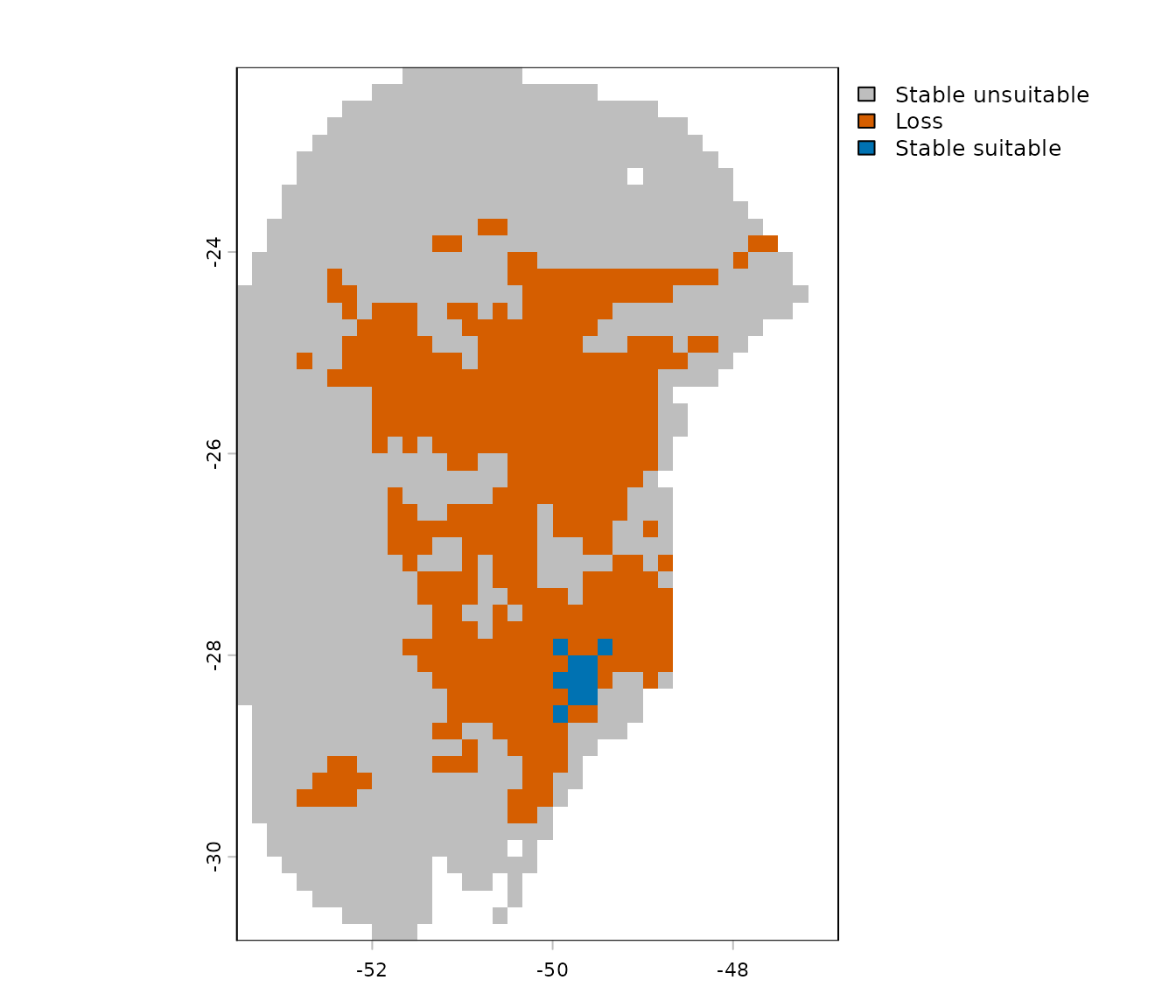
In this example, we are comparing current and future predictions, so
we set predicted_to = "future". If a comparison with past
predictions is needed, this argument should be set accordingly to ensure
that categories of change or stability are assigned correctly.
The prediction_changes() function is designed to compute
changes between single scenarios, meaning that the new scenario is
represented by one set of layers. If projections include multiple GCMs,
the function projection_changes() should be used instead.
For more details on projecting models and detecting changes with
summaries across multiple scenarios, see the vignette 6. Project Models to Multiple
Scenarios.